Snakemake—simple data processing
for researchers
Liang-Bo Wang
Taipei Bioinformatics Omnibus
2022-09-25
Esc or swipe up
with three fingers to exit
← → to navigate

Liang-Bo Wang, 2022-09-25.
Liang-Bo Wang
Taipei Bioinformatics Omnibus
2022-09-25
Esc or swipe up
with three fingers to exit
← → to navigate
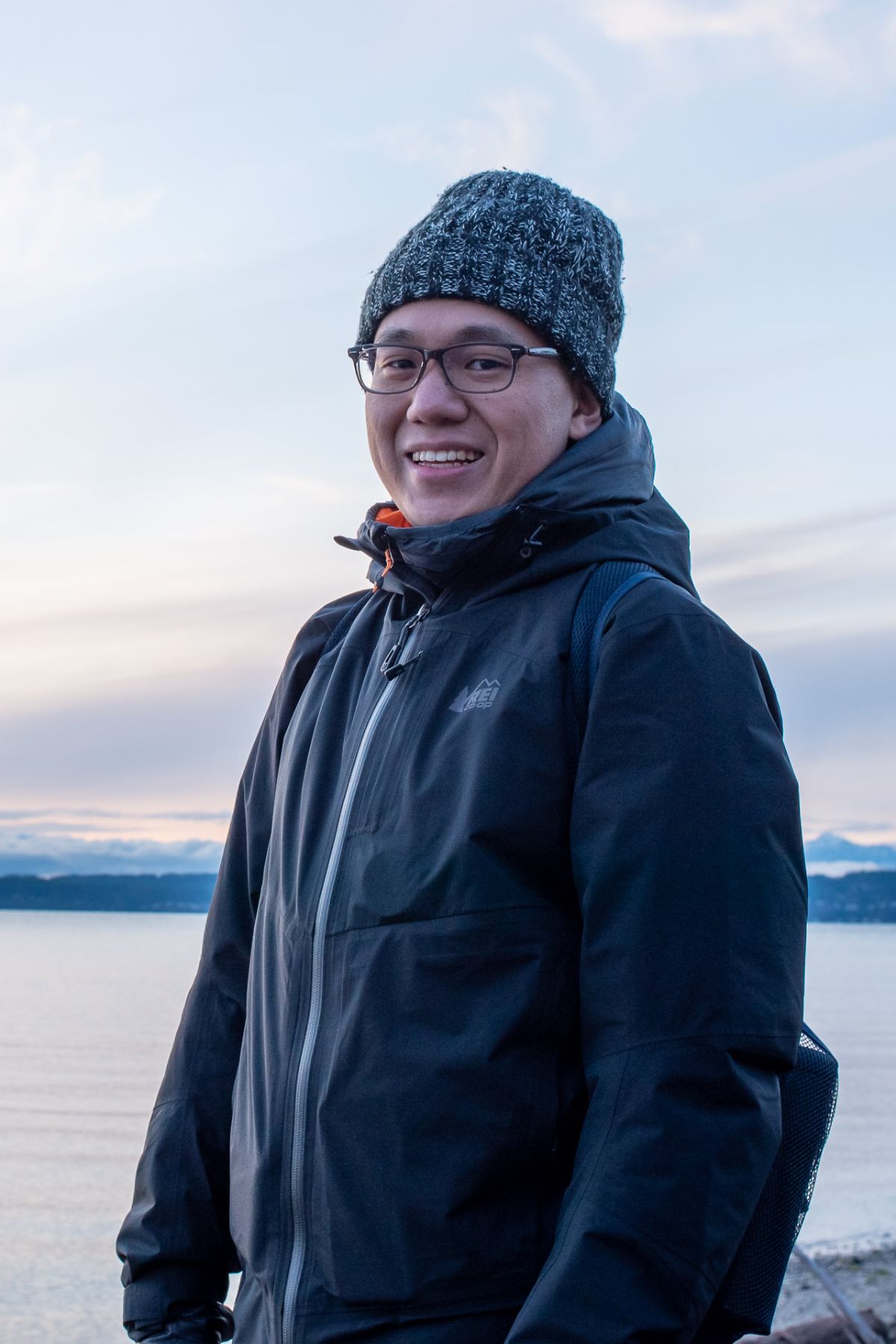



To process multiple samples on different computing platforms
To reproduce my analysis steps and results



 Data processing |
 Reproducible analysis |
|
|---|---|---|
| Inputs | Lots of samples | Mostly the same inputs |
| Run type | Once per sample | Multiple times |
| Workflow stability | Stable | Subject to change |
| Computing environment | Known and multiple Local, HPC, and cloud |
Known and limited* Mostly local |
There are quite a few completely opposite properties about these two scenarios, which are listed in the table.
For the reproducible analysis scenario, the computing environment is known and limited only when we try to reproduce it on our own machines. In my experience, having a clear and simple instructions is easier for other researchers to work with the analysis than say a fully automated and dockerized setup.
Snakemake is a Python-based workflow management tool.
A Snakemake workflow contains a set of rules:
Trace the workflow backwards. Snakemake starts with the rule that generates the final outputs:
The task dependency graph has the property as a directed acyclic graph (DAG). For Snakemake, such graph is constructed backwards from the final outputs, also known as the pull-based approach.
Reading the pipeline backwards can be a bit counterintuitive at first. My tip is to always start with the final outputs (usually at the end of the file) and trace the workflow backwards.
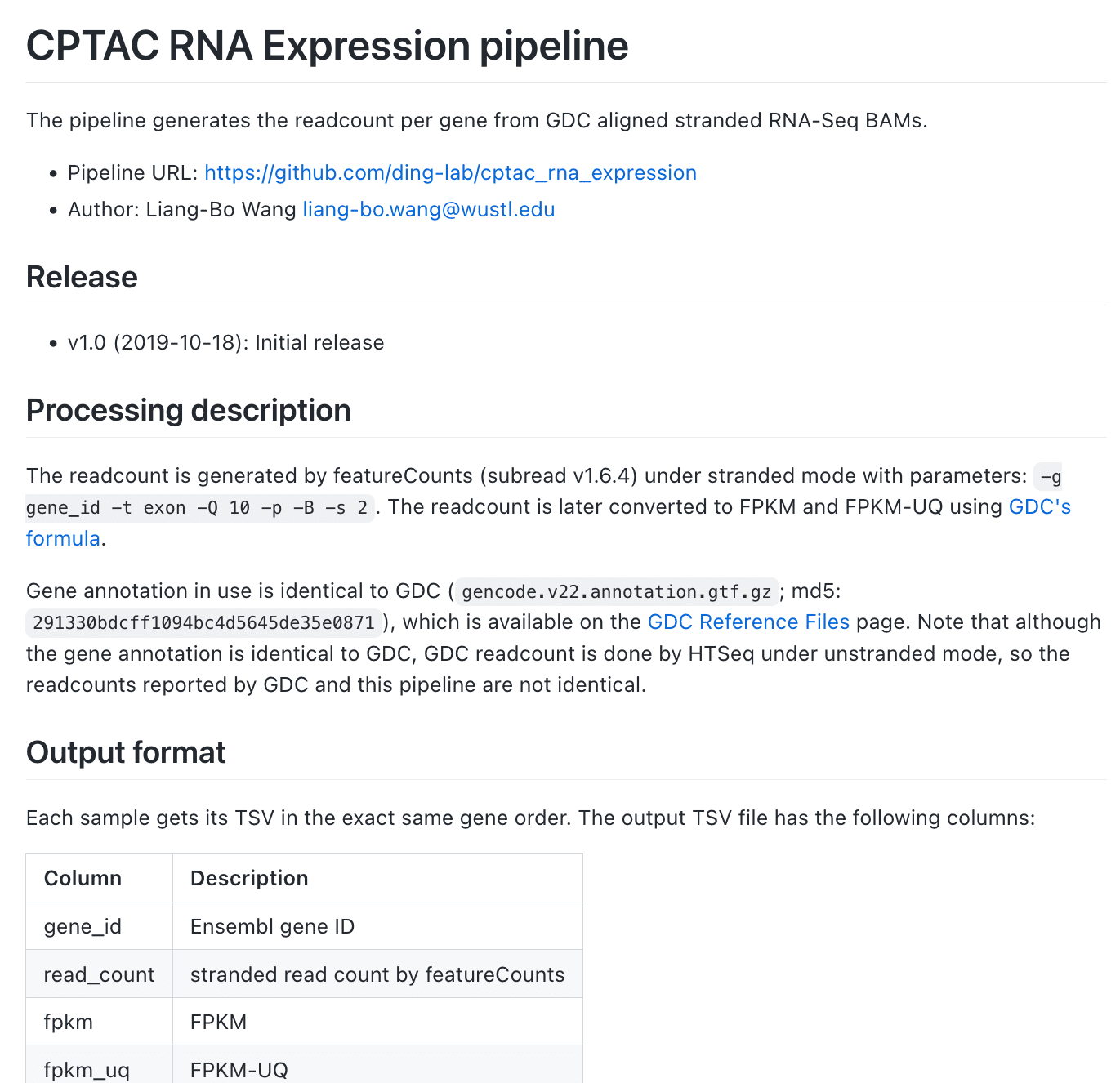

Calculate gene read counts from RNA-seq BAMs
Full pipeline here
I implemented two RNA-seq pipelines in Snakemake that are lossly based on GDC's mRNA analysis pipeline:
These pipelines were developed a while ago, when some of my computing environments were quite old and limited. So I took a relative conservative approach by using minimal features and very few dependencies (just the Python standard library). These pipelines can certainly be improved to better conform to the community's best practices. With the ubiquitous conda and Docker support and new Snakemake features, some possible improvements include:
{WORKFLOW_ROOT} with workflow.source_path()
rule readcount:
output: count_tsv='readcount/{sample}.tsv'
input: bam='inputs/{sample}.bam',
bai='inputs/{sample}.bam.bai'
log: 'logs/readcount/{sample}.log'
params: gtf=GENE_GTF_PTH
threads: 16
shell:
'featureCounts -T {threads} -a {params.gtf} ...'
'-o {output.count_tsv} {input.bam} 2> {log}'
{sample} is a wildcard whose value is determined by matching the output patterns
If you have wrote a Makefile before, snakemake rules share the same spirit as a Makefile's rule. And they run in a similar way.
{sample}input:, output:key='...' when there are multiple patterns. They are just Python strings and thus can be referred elsewhere by rules.myrule.output.key.shell:Check out the documentation for the rest parts of the rule.
Due to the way snakemake executes, it's easier to resolve the output first by realizing all the wildcard variables. Then expand the rest of the rule (inputs, parameters, and etc.)
For example, to generate readcount/my_sample_a.tsv using this task, {sample} expands to my_sample_a in the rest parts of the rule.
So Snakemake will look for the corresponding inputs and run the following shell commands:
featureCounts -T 16 -a /path/to/genes.gtf \
-o readcount/my_sample_a.tsv \
inputs/my_sample_a.bam \
2> logs/readcount/my_sample_a.log
snakemake all
To make the pipeline more extensible, use a run-all rule and a config file to process all samples of a batch.
As shown here, we obtain the list of samples to process from config file by
reading in the list from the file at config['sample_list'].
In this case, SAMPLES is a Python list of ["my_sample_a", "my_sample_b", ..., "my_sample_d"].
In the all rule, we use a Snakemake function expand() to generates all the output targets by "filling in" the wildcards of the output pattern using every value of SAMPLES.
When the simple string pattern matching is not enough, Snakemake also allows many parts of the rule to be determined by Python functions, including input, params, and resources.
These functions are known as input functions in Snakemake's documentation.
They make the rule more flexible.
As shown here, different samples may have their input files at different locations.
The same task will ask for more memory if it has been failed before.
unpack() is a helper function to automatically unpack the return value of the input function.
So a function can return multiple key-value pairs.
file_map_df is a pandas DataFrame object loaded from the tabular file specified by the config (config["file_map"]).
$ snakemake -j 32 readcount/my_sample_{a,b,c,d}.tsv
# featureCounts ... inputs/my_sample_a.bam
# featureCounts ... inputs/my_sample_b.bam
# ... (two jobs in parallel)
Alternatively, add a run-all rule to snakemake all:
with open(config['sample_list']) as f:
SAMPLES = f.read().splitlines()
rule all:
input: expand(rules.readcount.output.count_tsv, \
sample=SAMPLES)
Other powerful features that extend the wildcard file pattern matching:
rule readcount:
output: count_tsv='readcount/{sample}.tsv'
input: unpack(find_bam_path) # provides input.bam and input.bai
resources: # dynamic resource allocation
mem_mb=lambda wildcards, attempt: \
16000 + 16000 * (attempt - 1) # retry with more memory
...
def find_bam_path(wildcards):
# Retrieve local paths using the file map from config (a DataFrame)
bam_pth = file_map_df.at[wildcards.sample, "rna_genomic_bam_pth"]
return {'bam': bam_pth,
'bai': bam_pth + '.bai'}
I want to mention the dry run mode is very useful too (snakemake -n).

I actually had better experience to simply put together all the dependencies in one big fat Docker image or conda environment. This setup can live outside Snakemake, so it doesn't take care of pulling the docker image or creating the conda environment. This comes with some advantages:
However, this approach is against the best practice because it reduce the reusability of the rules and sometimes different rules have incompatible dependencies. One should consider switching to a rule-based environments/images later on.
pipeline_src/
├── scripts/...
└── Snakefile
batch_yyyy-mm-dd/ # from a template
├── config.json
├── sample.list
├── output/...
└── output_manifest.csv
# Under the batch folder
snakemake --configfile=config.json \
-s /pipeline_src/Snakefile \
-j 50 --attempt 3 \
--resources io_heavy=4 -- \
all
// config.json
{
"sample_list": "samples.list",
"file_map": "/path/to/file_map.csv",
"gtf": "/path/to/my_annotation.gtf",
...
}
# file_map.csv
sample,file_type,local_path,uuid,md5_checksum,...
sample_A,rna_bam,/path/to/sample_A.bam,5415b...
sample_B,rna_bam,/path/to/sample_B.bam,289c7...
# output_manifest.csv
sample,file_type,local_path,md5_checksum,...
sample_A,tsv,/path/to/readcount/sample_A.tsv,...
sample_B,tsv,/path/to/readcount/sample_B.tsv,...
Batch folders make the pipeline source code and the outputs live in different locations. Good for version tracking the pipeline and reusing the pipeline. We can also create a template folder for new batches. Config file is a natural solution to specify the batch-specific parameters.
Input file map is useful when the upstream files are tracked externally (say, in a database or some centralized system). It also retains the file metadata (e.g. checksums, UUID, experiment details, and other cross-references). For example, our lab maintains a file catalog to track all CPTAC data on GDC. It also tracks the files we have downloaded to the different storage systems in our lab. Thus, our example pipeline looks up this catalog to find the input BAM files.
Not sure if the file is the same? Verify its checksum!
There are quite a few possible reasons lead to a input change or corruption:
In general, it's a good idea to keep checksums all the time. Here is a snippet to generate MD5 checksums of everything under a folder in parallel and verify the content using the generated checksums:
fd --type file --exclude 'README*' --exclude '*CHECKSUMS' \
| sort \
| parallel --keep-order -j4 --bar 'md5sum {}' \
> MD5_CHECKSUMS
md5sum -c MD5_CHECKSUMS

09_xxx.Rmd)