王亮博 (亮亮)
Shared under CC 4.0 BY license
Esc to overview
← → to navigate


從哪裡開始,從哪裡結束。


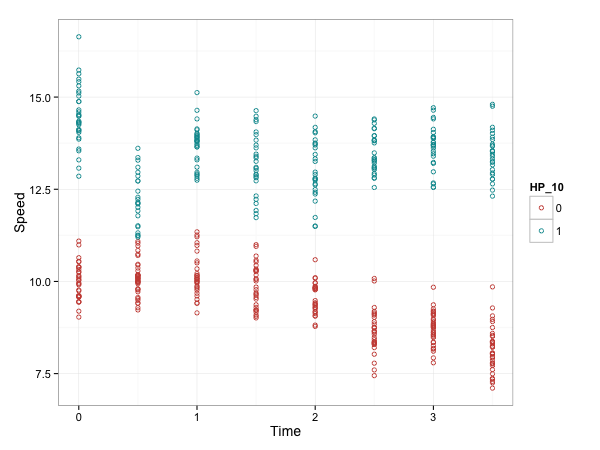
model.first <- lm(Speed ~ HP_10 + Time, data=df_sim)
# 更新模式
update(model.first, . ~ . - HP_10)
update(model.first, . ~ . + HP_10:Time) # interaction
這個 Python 大概也做得到,沒什麼 ˊ_>ˋ
mod_trans <- lm(
log(Speed) ~ HP_10 * Time + I(Time^2), data=df_sim)
summary(mod_trans)
# Residuals:
# Min 1Q Median 3Q Max
# -0.185972 -0.043402 0.002693 0.042158 0.211709
# Coefficients:
# Estimate Std. Error t value Pr(>|t|)
# (Intercept) 2.341137 0.008649 270.670 < 2e-16 ***
# HP_101 0.258697 0.010874 23.790 < 2e-16 ***
# Time -0.052432 0.009884 -5.305 1.74e-07 ***
# I(Time^2) -0.003214 0.002621 -1.226 0.221
# HP_101:Time 0.058656 0.005207 11.264 < 2e-16 ***
# ---
# Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
# Residual standard error: 0.066 on 473 degrees of freedom
# Multiple R-squared: 0.8916, Adjusted R-squared: 0.8907
# F-statistic: 972.7 on 4 and 473 DF, p-value: < 2.2e-16
model.full <- lm(Speed ~ HP_10 * Time, data=df_sim)
model.hi <- update(
model.full,
. ~ . + HP_10 : I(Time^2), # try `HP_10 * I(Time^2)`
data=df_sim
)
anova(model.full, model.hi)
summary(model.hi)
anova(...)
# Analysis of Variance Table
#
# Model 1: Speed ~ HP_10 * Time
# Model 2: Speed ~ HP_10 + Time + HP_10:Time + HP_10:I(Time^2)
# Res.Df RSS Df Sum of Sq F Pr(>F)
# 1 474 269.43
# 2 472 242.26 2 27.175 26.473 1.269e-11 ***
summary(...)
# Coefficients:
# Estimate Std. Error t value Pr(>|t|)
# (Intercept) 10.09593 0.10485 96.289 < 2e-16 ***
# HP_101 3.87381 0.15231 25.433 < 2e-16 ***
# Time 0.06238 0.14318 0.436 0.663
# HP_101:Time -0.94099 0.20679 -4.551 6.81e-06 ***
# HP_100:I(Time^2) -0.18324 0.03914 -4.681 3.73e-06 ***
# HP_101:I(Time^2) 0.23082 0.04143 5.571 4.27e-08 ***
geom_smooth() 可以用
g + geom_point(shape=1) +
geom_smooth(method=lm, se=TRUE)
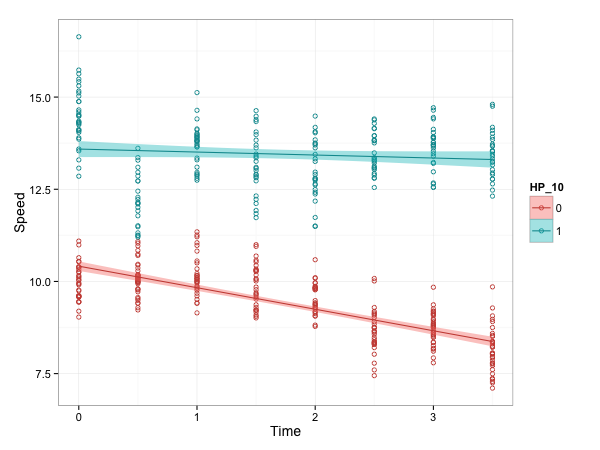
y ~ x使用 stat_smooth()
stat_smooth(
method="lm", formula = y ~ x + I(x^2),
se=FALSE) # use y and x in formula
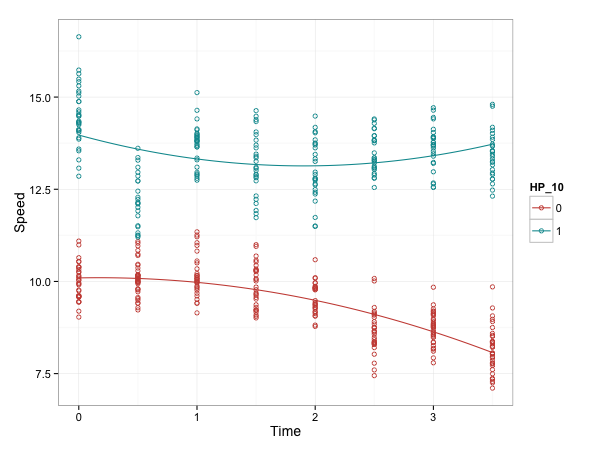
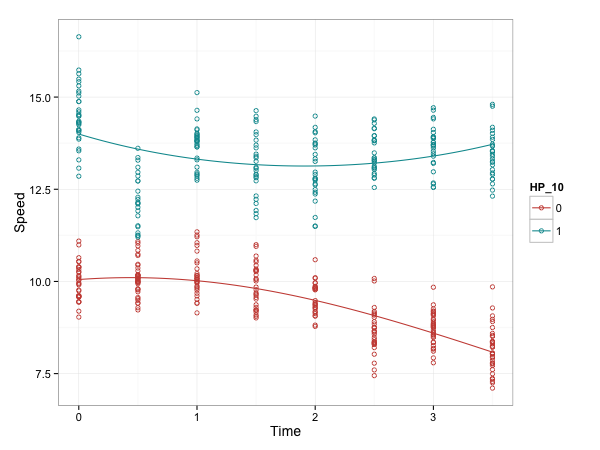
g + geom_point(shape=1) + stat_smooth(
method="glm", family=gaussian(link="inverse"),
formula = y ~ x + I(x^2), se=FALSE)

控制不同城市後,肺癌在有無吸煙人的發生率是不是相同?
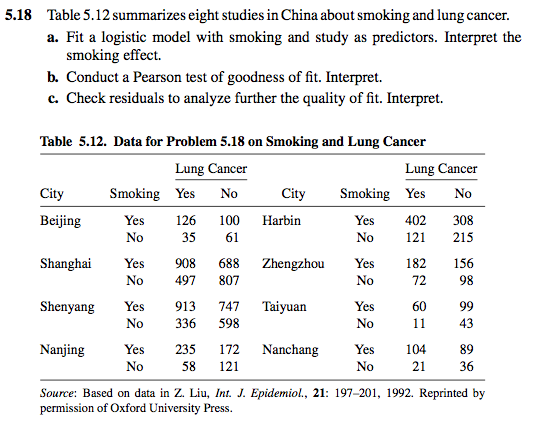
china_smoke <- read.table("chismoke.dat", header=TRUE)
# chinasmoke.dat
# City Smoker Cancer Count
# Beijing Yes Yes 126
# Beijing Yes No 100
# Beijing No Yes 35
# Beijing No No 61
# ...
tab_cs <- xtabs(Count ~ Cancer + Smoker + City, china_smoke)
ftab_cs <- ftable(tab_cs, row.vars=c("City", "Smoker"))
ftab_cs
# Cancer No Yes
# City Smoker
# Beijing No 61 35
# Yes 100 126
# Harbin No 215 121
# Yes 308 402
# Nanchang No 36 21
# Yes 89 104
# Nanjing No 121 58
# Yes 172 235
# ...
library(reshape2)
d_cs <- dcast(china_smoke, City + Smoker ~ Cancer, value.var="Count")
model <- glm(cbind(Yes, No) ~ Smoker + City, data=d_cs, family=binomial)
summary(model)
## to LaTeX
library(xtable)
xtab <- xtable(model, digits=3)
print(xtab, booktabs = TRUE)
print(xtable(d_cs[, c(-5)]), booktabs = TRUE)
library(memisc)
toLatex(ftab_cs, useDcolumn=FALSE)